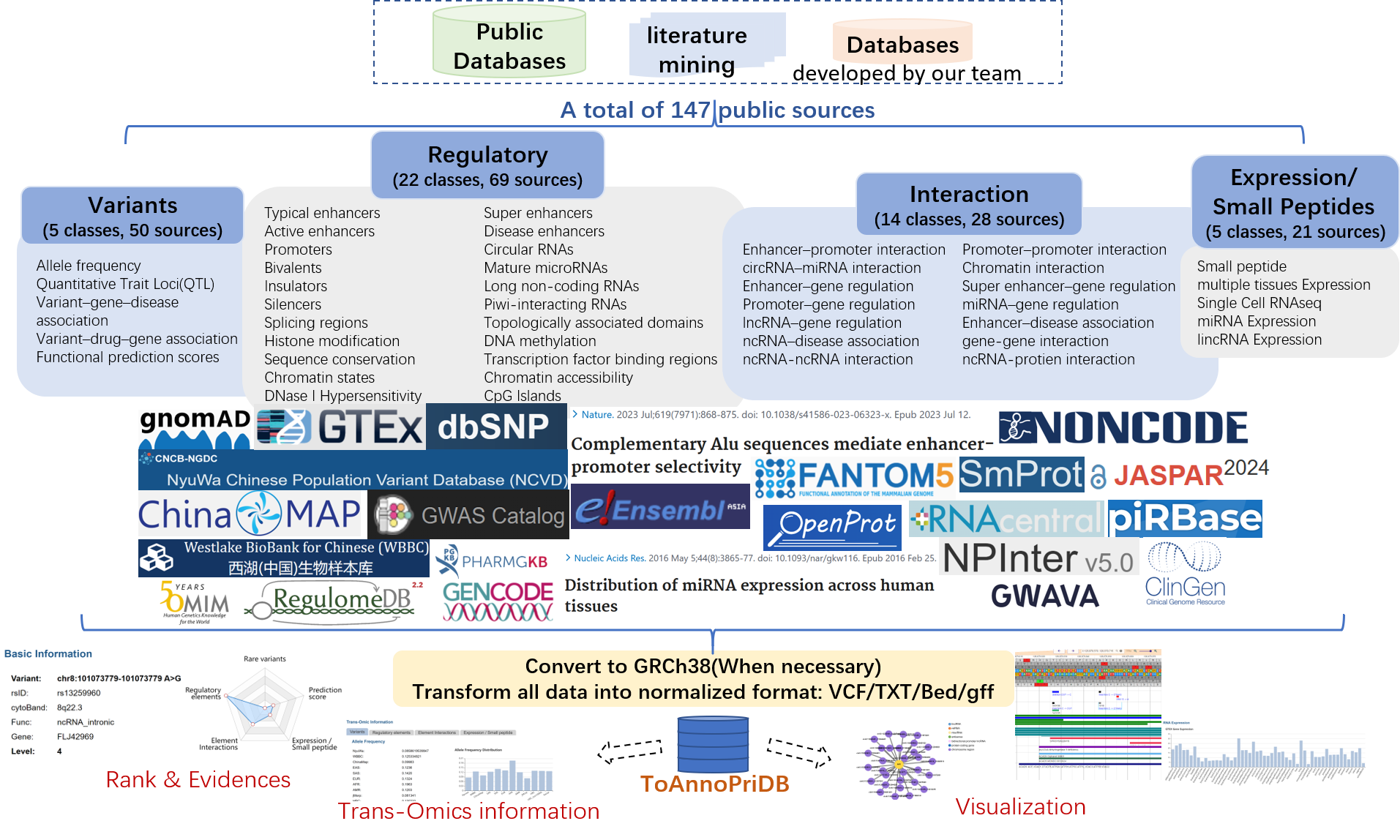
Non-coding variants can affect gene expression and regulation through various mechanisms, including altering transcription factor binding, histone modifications, biomolecule interactions, and chromatin accessibility, among others. Therefore, gaining a comprehensive understanding of the functional significance of mutations may require considering multi-omics information rather than relying on a single aspect. Thus, based on 147 public sources including our own databases (NyuWa, NONCODE, NPInter, piRBase, SmProt, LncVar), we developed the TOAnnoPriDB database with the aim of annotating and ranking non-coding variants from a multi-omics perspective.
TOAnnoPriDB includes allele frequency information from 15 different populations, particularly population frequency information for 237,373,360 variants from 25,432 individuals of Chinese descent, over 12,053,317 experimentally tested non-coding variants with expression-modulating potentials, and 15,514,528 disease-associated variant entries, covering 46 types of multi-omics annotations including population frequency, functional prediction, disease association, regulation, interaction, and translational expression.
The objective of TOAnnoPriDB is to assist researchers in selecting and prioritizing variants for further study, providing more targeted directions for investigation and validation, and exploring the associations between non-coding variants and human diseases.