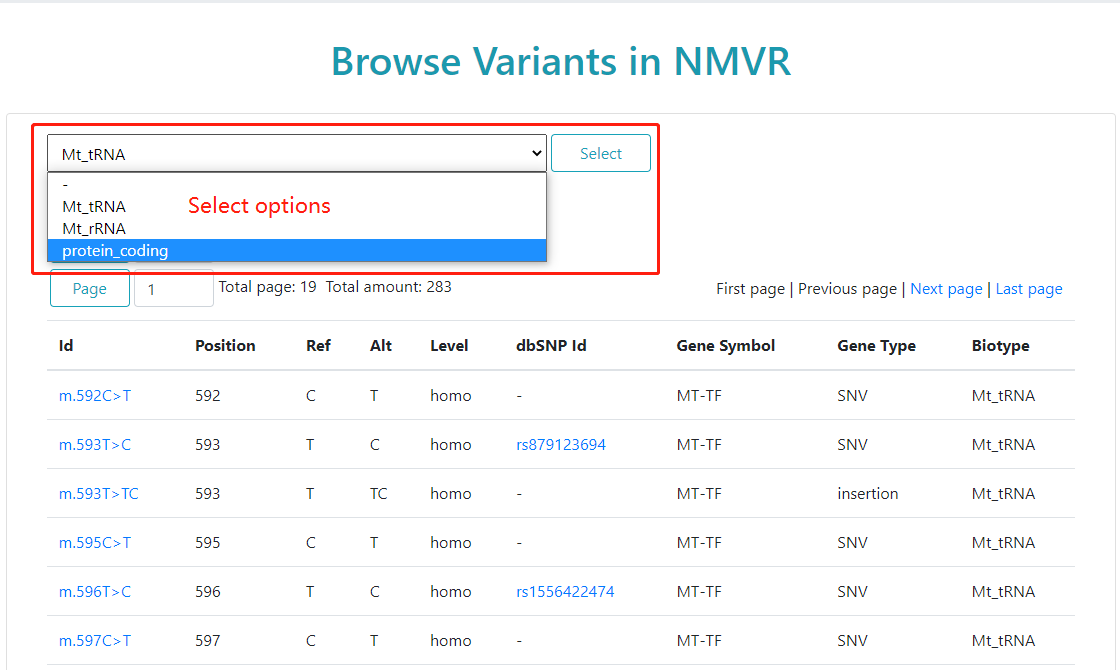
Users can browse NMVR by mtDNA feature types.
The select options include Mt_tRNA, Mt_rRNA, protein_coding,
and "-" for non-gene (D-loop region and intergenic regions).
Users can click on the Variant Id to go to the details page
of mtDNA variants in NMVR. Some mtDNA variants are annotated to RS in dbSNP,
and NMVR provides links to these variants for further information.
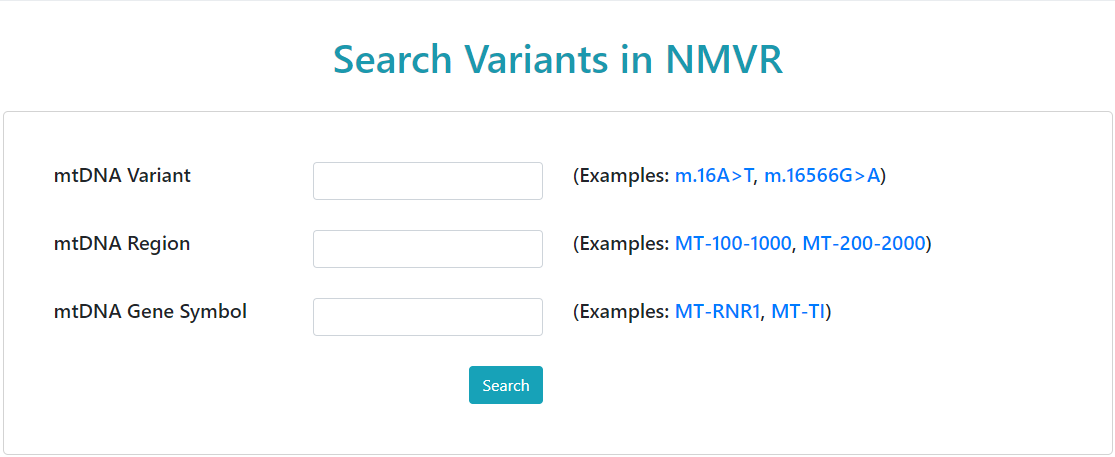
Users can search NMVR by entering the keywords like mtDNA variant,
gene symbol, and mtDNA region.
The keyword format:
- mtDNA Variant:
m.4696T>C, m for mitochondria, 4696 for mtDNA position,
T>C for referece base(s) to alternative base(s).
- mtDNA Region:
MT-4000-5000, MT for mitochondria, 4000 for the region start,
5000 for the region end. The delimiter must be "-".
- mtDNA Gene Symbol:
MT-ND2, MT for mitochondria.
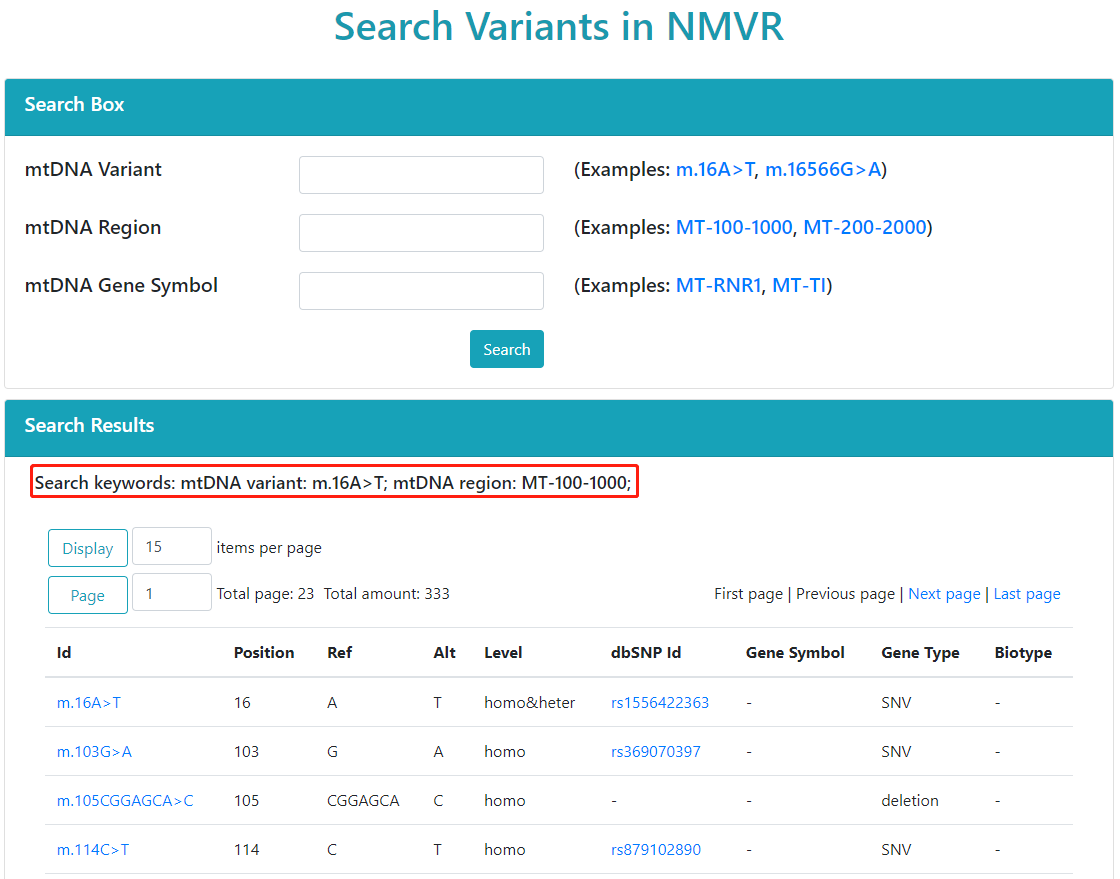
The search results page is divided into two parts.
The upper part is the search box, the user can re-enter keywords to search.
The lower part shows the results of the search.
Users can enter multiple different keywords at the same time。
Users can click on the Variant Id to go to the details page
of mtDNA variants in NMVR. Some mtDNA variants are annotated to RS in dbSNP,
and NMVR provides links to these variants for further information.
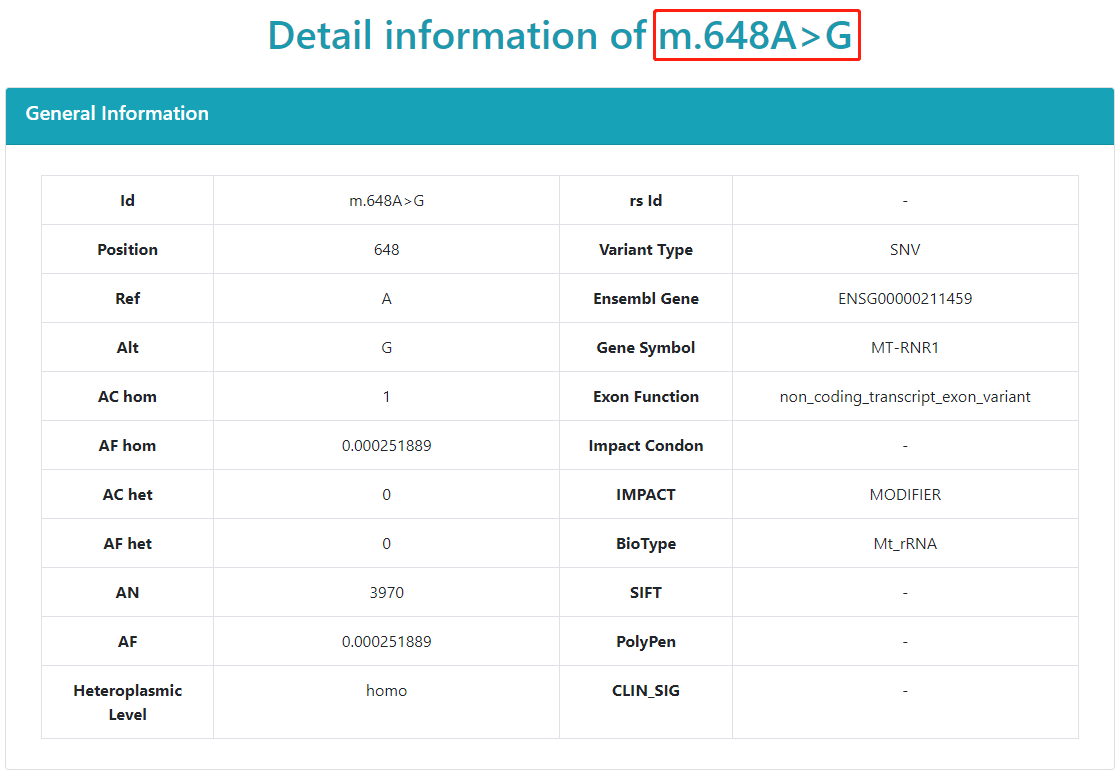
3. The detail information of mtDNA variants
The mtDNA variant Id is given in the title.
The information of the mtDNA variants in NMVR is
displayed in the General Information panel.
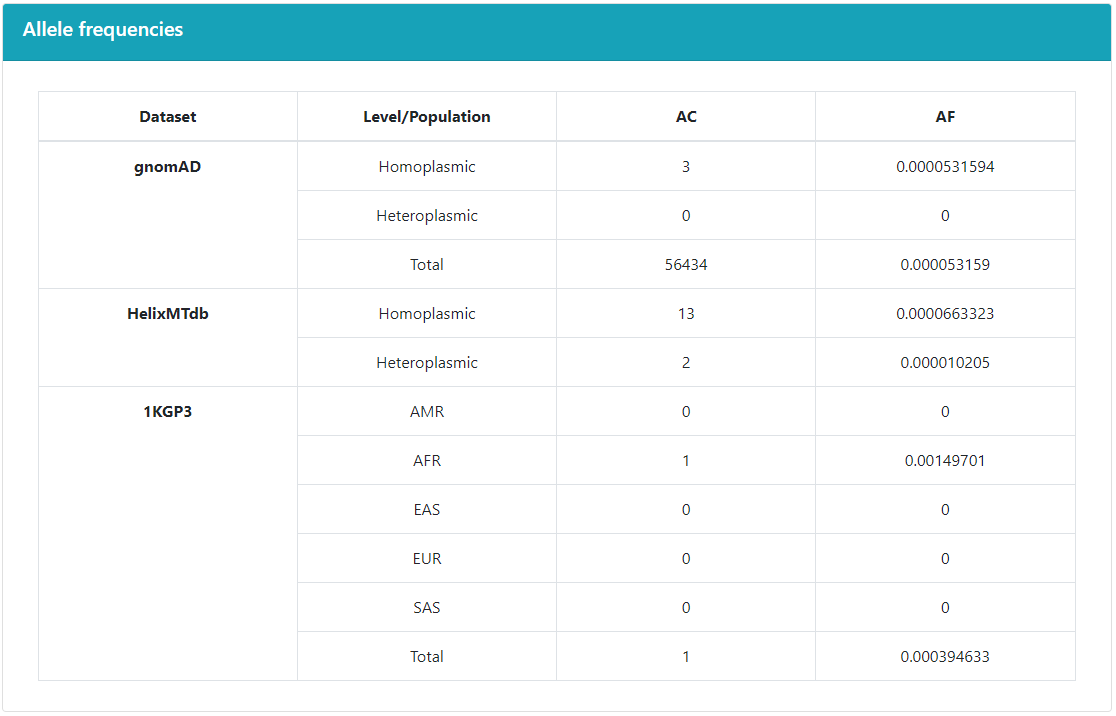
The Allele frequencies panel shows the information
in other databases.
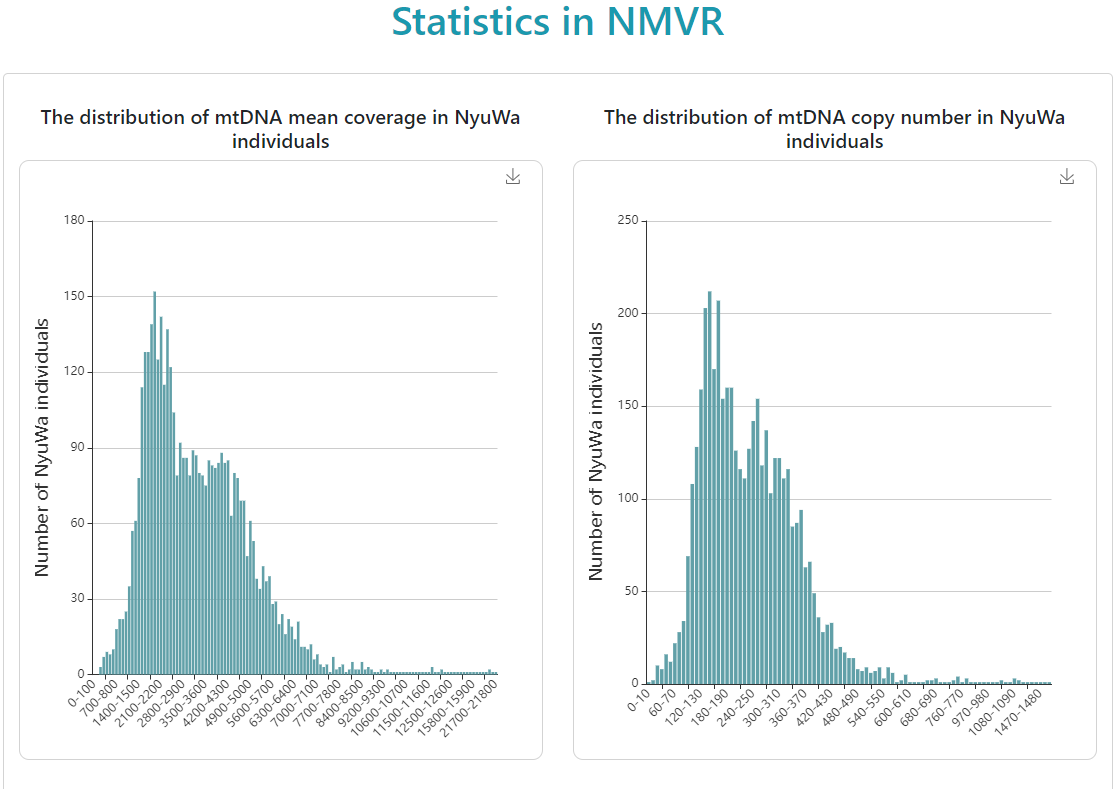
Show the statistical results from NMVR.
-
Distribution of mtDNA mean coverage in NyuWa samples.
-
Distribution of mtDNA copy number in NyuWa samples.
-
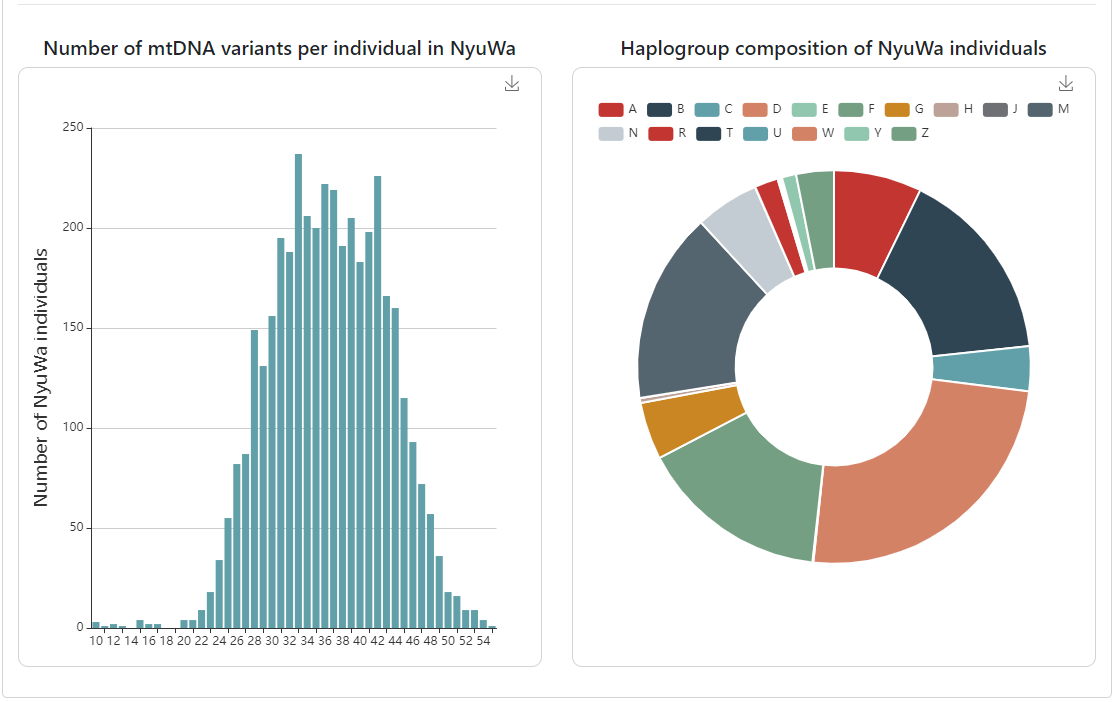
Number of high-quality mtDNA variants per individual
in NyuWa.
-
The macrohaplogroup (first letter of the haplogroup
reported by haplogrep) composition of NyuWa samples.